2026-02-25 12:33:48
Crosslisted article(s) found for cs.LG. https://arxiv.org/list/cs.LG/new
[3/3]:
- Functional Continuous Decomposition
Teymur Aghayev
https://arxiv.org/abs/2602.20857 https://mastoxiv.page/@arXiv_eessSP_bot/116130499236089653
- SpatiaLQA: A Benchmark for Evaluating Spatial Logical Reasoning in Vision-Language Models
Xie, Zhang, Shan, Zhu, Tang, Wei, Song, Wan, Song
https://arxiv.org/abs/2602.20901 https://mastoxiv.page/@arXiv_csCV_bot/116130845273808954
- Some Simple Economics of AGI
Christian Catalini, Xiang Hui, Jane Wu
https://arxiv.org/abs/2602.20946 https://mastoxiv.page/@arXiv_econGN_bot/116130470423837005
- Multimodal MRI Report Findings Supervised Brain Lesion Segmentation with Substructures
Yubin Ge, Yongsong Huang, Xiaofeng Liu
https://arxiv.org/abs/2602.20994 https://mastoxiv.page/@arXiv_eessIV_bot/116130212832138624
- MIP Candy: A Modular PyTorch Framework for Medical Image Processing
Tianhao Fu, Yucheng Chen
https://arxiv.org/abs/2602.21033 https://mastoxiv.page/@arXiv_csCV_bot/116130864279556063
- Empirically Calibrated Conditional Independence Tests
Milleno Pan, Antoine de Mathelin, Wesley Tansey
https://arxiv.org/abs/2602.21036 https://mastoxiv.page/@arXiv_statME_bot/116130690605113562
- Is Multi-Distribution Learning as Easy as PAC Learning: Sharp Rates with Bounded Label Noise
Rafael Hanashiro, Abhishek Shetty, Patrick Jaillet
https://arxiv.org/abs/2602.21039 https://mastoxiv.page/@arXiv_statML_bot/116130572661848449
- Position-Aware Sequential Attention for Accurate Next Item Recommendations
Timur Nabiev, Evgeny Frolov
https://arxiv.org/abs/2602.21052 https://mastoxiv.page/@arXiv_csIR_bot/116130263323086316
- Motivation is Something You Need
Mehdi Acheli, Walid Gaaloul
https://arxiv.org/abs/2602.21064 https://mastoxiv.page/@arXiv_csAI_bot/116130680774678580
- An Enhanced Projection Pursuit Tree Classifier with Visual Methods for Assessing Algorithmic Impr...
Natalia da Silva, Dianne Cook, Eun-Kyung Lee
https://arxiv.org/abs/2602.21130 https://mastoxiv.page/@arXiv_statML_bot/116130610674573081
- Complexity of Classical Acceleration for $\ell_1$-Regularized PageRank
Kimon Fountoulakis, David Mart\'inez-Rubio
https://arxiv.org/abs/2602.21138 https://mastoxiv.page/@arXiv_mathOC_bot/116130547076073836
- LUMEN: Longitudinal Multi-Modal Radiology Model for Prognosis and Diagnosis
Jiang, Yang, Nath, Parida, Kulkarni, Xu, Xu, Anwar, Roth, Linguraru
https://arxiv.org/abs/2602.21142 https://mastoxiv.page/@arXiv_csCV_bot/116130871488694585
- A Benchmark for Deep Information Synthesis
Debjit Paul, et al.
https://arxiv.org/abs/2602.21143 https://mastoxiv.page/@arXiv_csAI_bot/116130692571594706
- Scaling State-Space Models on Multiple GPUs with Tensor Parallelism
Anurag Dutt, Nimit Shah, Hazem Masarani, Anshul Gandhi
https://arxiv.org/abs/2602.21144 https://mastoxiv.page/@arXiv_csDC_bot/116130520888343997
- Not Just How Much, But Where: Decomposing Epistemic Uncertainty into Per-Class Contributions
Mame Diarra Toure, David A. Stephens
https://arxiv.org/abs/2602.21160 https://mastoxiv.page/@arXiv_statML_bot/116130618512594211
- Aletheia tackles FirstProof autonomously
Tony Feng, et al.
https://arxiv.org/abs/2602.21201 https://mastoxiv.page/@arXiv_csAI_bot/116130705679345625
- Squint: Fast Visual Reinforcement Learning for Sim-to-Real Robotics
Abdulaziz Almuzairee, Henrik I. Christensen
https://arxiv.org/abs/2602.21203 https://mastoxiv.page/@arXiv_csRO_bot/116130765974498223
toXiv_bot_toot
2025-12-14 19:25:37
It was approaching time to renew my Global Entry Membership. One of the questions required for renewal was to list the countries I’ve been to since 2020.
Each country visited had to be selected from a list. The UI was excellent. I could type a substring and the options would be limited to countries containing that substring ✅. I couldn’t find Turkey. Searched Tu, scrolled the unfiltered list no Turkey.
WHY?
2026-01-15 23:20:50
Formation of Substructure in Luminous Submillimeter Galaxies (FOSSILS) - Evidence of Multiple Pathways to Trigger #Starbursts in Luminous Submillimeter Galaxies: https://iopscience.iop.org/article/10.3847/1538-4357/ae157e -> Multiple Origins Behind The Extreme Star Formation In “Monster Galaxies”: https://alma-telescope.jp/en/news/press/multipleorigins-202601.html
2026-01-14 01:45:53
Sources: Apple and Qualcomm are scrambling to secure glass cloth fiber, used in chip substrates and PCBs, as the AI boom boosts demand for the component (Nikkei Asia)
https://asia.nikkei.com/business/technology/tech-…
2026-01-15 13:00:14
2026-02-07 03:00:04
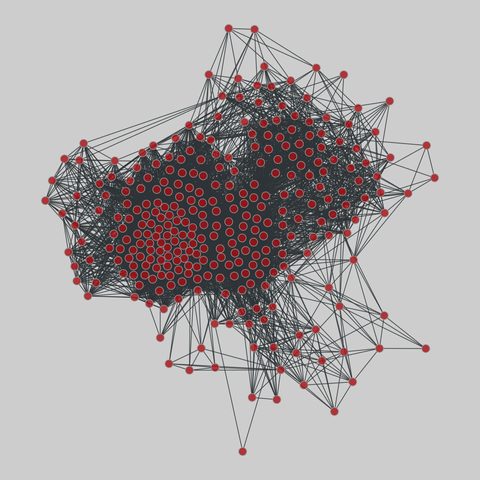
malaria_genes: Malaria var DBLa HVR networks
Networks of recombinant antigen genes from the human malaria parasite P. falciparum. Each of the 9 networks shares the same set of vertices but has different edges, corresponding to the 9 highly variable regions (HVRs) in the DBLa domain of the var protein. Nodes are var genes, and two genes are connected if they share a substring whose length is statistically significant. Metadata includes two types of node labels, both based on sequence st…
2025-12-12 12:04:13
Two-dimensional semiconductor-based active array for high-fidelity spatiotemporal monitoring of neural activities (in mice) #BCI
2026-01-02 14:00:04
malaria_genes: Malaria var DBLa HVR networks
Networks of recombinant antigen genes from the human malaria parasite P. falciparum. Each of the 9 networks shares the same set of vertices but has different edges, corresponding to the 9 highly variable regions (HVRs) in the DBLa domain of the var protein. Nodes are var genes, and two genes are connected if they share a substring whose length is statistically significant. Metadata includes two types of node labels, both based on sequence st…
2025-11-26 19:00:03
malaria_genes: Malaria var DBLa HVR networks
Networks of recombinant antigen genes from the human malaria parasite P. falciparum. Each of the 9 networks shares the same set of vertices but has different edges, corresponding to the 9 highly variable regions (HVRs) in the DBLa domain of the var protein. Nodes are var genes, and two genes are connected if they share a substring whose length is statistically significant. Metadata includes two types of node labels, both based on sequence st…
2025-12-26 13:00:04
malaria_genes: Malaria var DBLa HVR networks
Networks of recombinant antigen genes from the human malaria parasite P. falciparum. Each of the 9 networks shares the same set of vertices but has different edges, corresponding to the 9 highly variable regions (HVRs) in the DBLa domain of the var protein. Nodes are var genes, and two genes are connected if they share a substring whose length is statistically significant. Metadata includes two types of node labels, both based on sequence st…